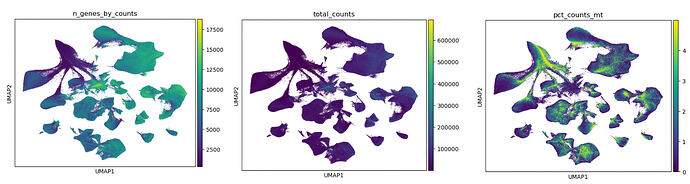
Here are the plots. I was wondering if I should at pct_counts_mt as a categorical covariate in scVI? Additionally n_genes_by_counts does show lower gene detection in the problematic quadrant of the UMAP however, it is only “removed“ when I do data = adata[adata.obs[‘n_genes_by_counts’] >= 2000, :] in the same latent space, which is really stringent.