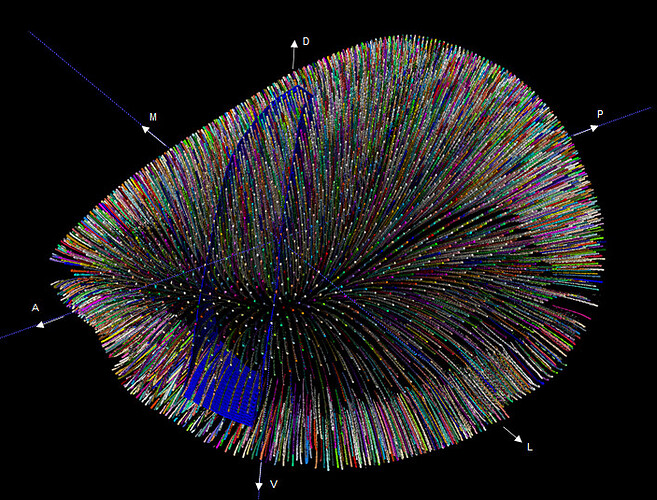
We are often asked: “Why do the boundaries on the CCF look so odd for some cortical areas?”.
Here, they are referring to the coronal views of the Allen Mouse Brain Common Coordinate Framework (CCF) in the 2D atlas viewer. The short answer is that in those places the cortical surface is tilted from the horizontal such that the resulting oblique view gives the impression that the cortical boundary is not perpendicular to the cortical surface.
Establishing a curved cortical framework
The longer answer starts with considering the isocortex: a 3D sheet organized into layers where connections between the layers are typically perpendicular to the surface in a columnar or radial organization. The curvature of the isocortex makes it difficult to visualize along this perpendicular dimension, as one “column” or “radial” can cut through a variable number of coronal/sagittal planes depending on where it is located. To enable the visualization and integration of data along the “columns”, a curved cortical framework was developed (Wang et al, 2020). A set of streamlines were computed that approximates the path of the radials through the cortical sheet based on the geometry of the cortex.
The curved cortical framework was then used to project data along the streamlines to create surface views of the CCF average template (grayscale image below) and other reference data to guide the manual annotation of the 2D views (pink overlay).
From the 2D parcellation, a 3D parcellation was generated by assigning to each entire streamline the same cortical area that was assigned to it on the surface, resulting in a 3D cortical parcellation (insert) that is consistent with reference data and the curved cortical framework.
Case study: the MOp and SSp border
The boundary between MOp (primary motor area) and SSp (primary somatosensory area) is primarily defined by the presence/absence of layer 4 containing somatotopically organized domains. These domains are clearly visible in the average template and surface views, and along with supporting data (see end of the post) enables straightforward drawing of a 2D boundary on a surface view.
This boundary corresponds to a set of streamlines which together forms a curved interface representing a 3D boundary between MOp and SSp. By construction, this 3D boundary is consistent with both layer 4 information and the curvature of the cortex.
Let’s take a look again at the coronal section at the top of the post. At first glance, the appearance of a almost vertical MOp/SSp border seems counterintuitive to the claim we just made above. In the figures below, we superimpose some of the streamlines onto the section to help illustrate what is going on.
The first row of the figure shows a sagittal (A) and coronal (B) section from CCFv3 with the annotated structures shown as blue outlines. A regular sub-sample of the streamlines are dilated and superimposed with randomly assigned colors. The crosshairs are centered on the same streamline (red) in all views.
A streamline (just like real apical dendrites on a cell) will typically intersect multiple coronal/sagittal planes. Conversely, in any single plane, only a segment of the streamline will intersect the plane. The size of the segment varies along the cortex, where the longer the segment the more a streamline is “parallel” to the viewing plane. The orientation of the segment follows the shortest path from pia to white matter surface and tends to be perpendicular to the surfaces.
At the crosshair location, the pia surface is titled 23 degree down and as such on the vertical coronal plane the MOp/SSp border is composed of segments of many different streamlines as seen in the zoomed in sagittal © and coronal (D) views. Due to the tilt, the vertical appearance of the border is actually what we would typically think of as horizontal components of the areal border showing through.
You can see this effect if you cut up an orange in an oblique (off-vertical) angle.
After we rotate the brain by 23 degrees so that the pia surface is now lying horizontally at the crosshair location, we can generate a new parasagittal (E) and paracoronal (F) section. The new paracoronal section is now approximately parallel to the streamlines resulting in larger streamline segments being visible in the overlay (H) and the MOp/SSp boundary now appears as a 2D perpendicular line to the surface contour.
With the rotation, a larger portion of the cortex is now approximately horizontal, so the same “straightening” effect can now be seen in the subsequent sections 0.4, 0.8 and 1.2 mm to the posterior in the figure below (top row: before rotation, bottom row: after rotation) demonstrating the intuition that cortical area borders should be approximately perpendicular to the cortical surface has been faithfully retained in the CCFv3 and is fully consistent in 3D.
While we are using MOp/SSp as a case study for this post, delineation of all cortical areas in CCFv3 followed the same principle and additionally validated also in 3D volumetric views by our neuroanatomy team.
Supporting reference data
MOp/SSp border
- Scnna1a-Tg3-Cre transgene expression (enriched expression in layer 4)
- Nr5a1-Cre transgene expression (enriched expression in layer 4)
- Rorb expression shows a narrow band at the transition from SSp/MOp but exists in MOp, while Scnna1a and Nr5a1 expression stops at the border. CCFv3 follows the convention of Allen Reference Atlas (Dong, 2008) where layer 4 is absent in MOp and hence Scnna1a and Nr5a1 expression is used but not Rorb.
MOp/MOs border
- Grp-Cre-KH288 transgene expression (enriched expression in MOs)
- VAL axonal projection data , top view (projection to MOp from VAL)
SSp/SSs border
- Nr5a1-Cre transgene expression (enriched in layer 4, weak in SSs)
- Rorb-IRES2-Cre transgene expression (enriched in layer 4, weak in SSs)
Isocortical layers
- Calb1-T2A-dgCre transgene expression (enriched in layer 2/3)
- Nr5a1-Cre transgene expression (enriched in layer 4)
- Rbp4-Cre_KL100 transgene expression (enriched in layer 5)
- Ntsr1-Cre_GN220 transgene expression (enriched in layer 6a)
- Ctgf-T2A-dgCre transgene expression (enriched in layer 6b)
View and access all the transgene expression data used for delineating CCFv3.
See registered transgene expression and axonal projection data overlaid with CCFv3 structure outlines:
See streamlines overlaid on CCFv3 average template and structures:
Feedback
Got questions or related insights and data to share? Post a note or comment below.